IMProv - Mass Spec Studio Configuration Tutorial
This tutorial presents the msstudio wizard to gather the data files and driver scripts needed to perform a MPI based IMP modeling job run. We cover how this can be prepared using the IMProv wizard in MassSpecStudio.
PRC2 example project (download):
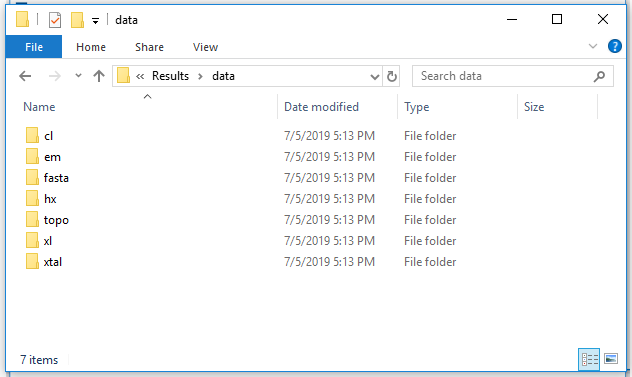
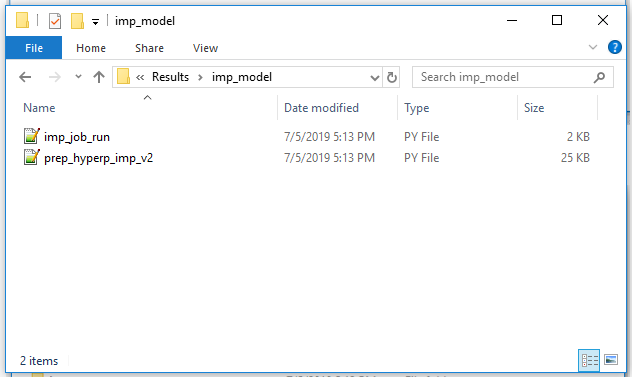
The data folder contains various artifacts used to inform the integrative modeling. The imp_model folder contains the driver python script and example yaml configuration for running the IMP modeling job.
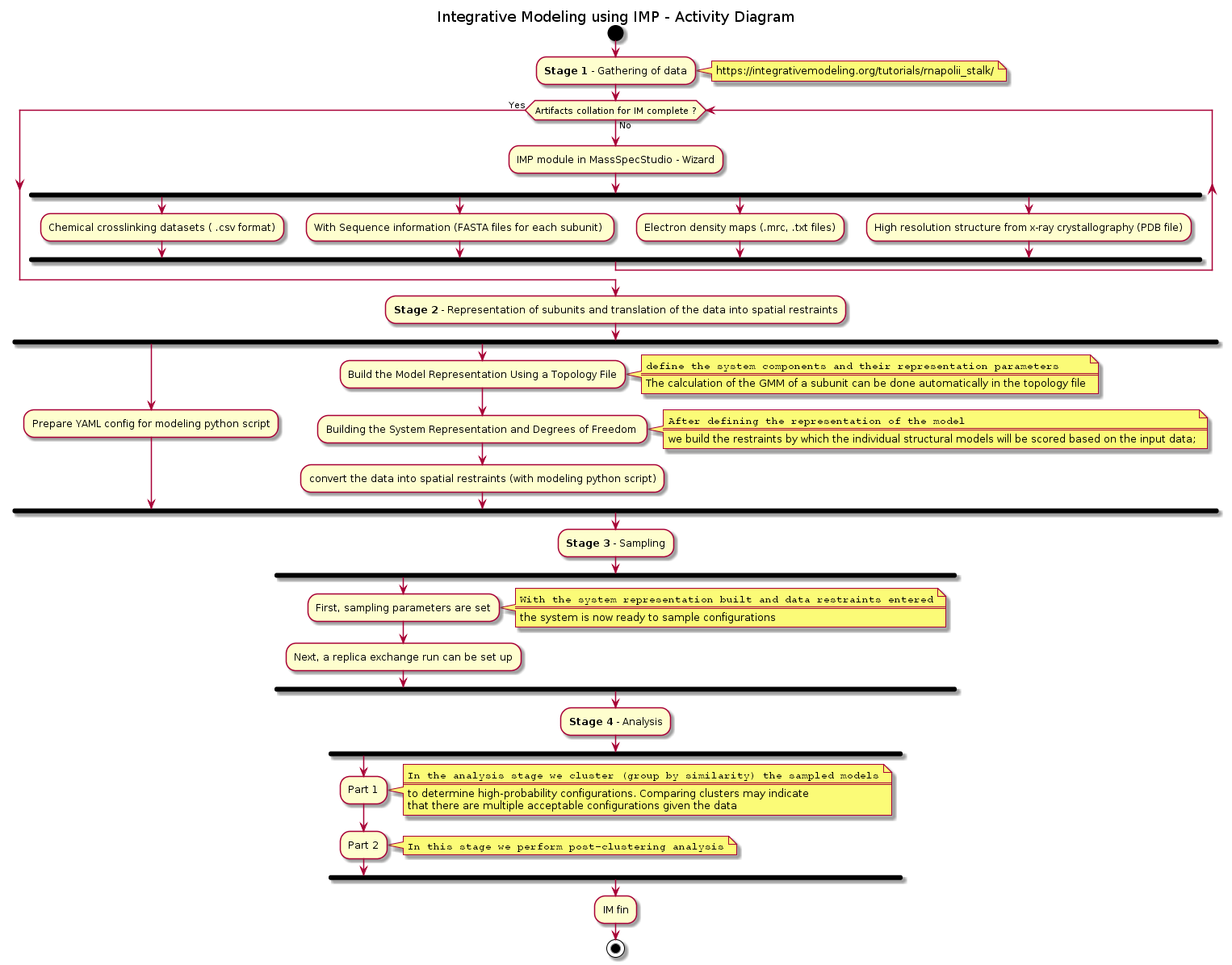
We are able to prepare the data files and driver scripts given above using Mass Spec Studio's IMProv wizard. The activity diagram that follows illustrates the various steps involved in running the modeling pipeline. This is provided here in order to set the context for the pre-requisite steps that we need to perform prior to deployment for this modeling job run.
MassSpecStudio is used to configure the artifacts that are subsequently involved in the following activity diagram:
Prepare a new IMProv modeling project with Mass Spec Studio
In order to pull together the various data files and prepare the artifacts of a IMProv modeling bundle. Mass Spec Studio provides a wizard that guides the user through the selection of the data files and setup of the various configuration settings required by the IMP modeling run.
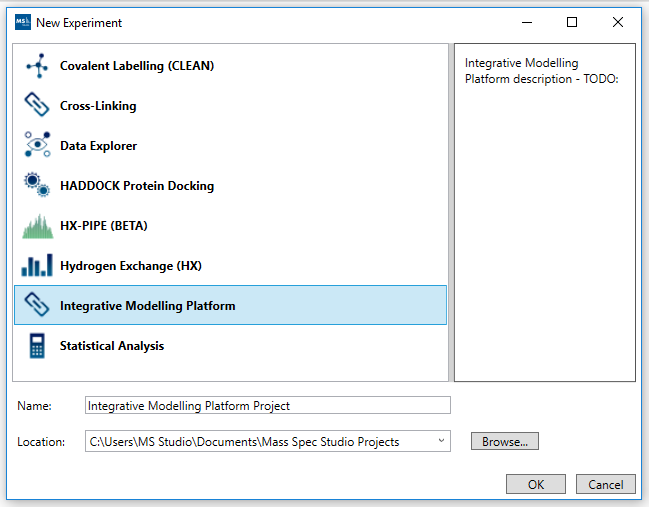
1. Project initialization:
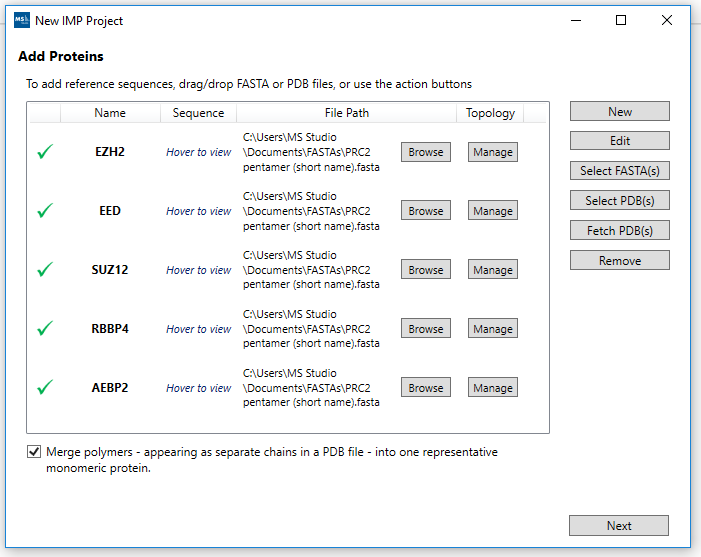
2. Add proteins:
Using the Add Proteins wizard screen. Select the FASTA or PDB files to add reference sequences. This will then show the Name and give the opportunity to customize the Topology by clicking the Manage button under the Topology column for the row with a protein name.
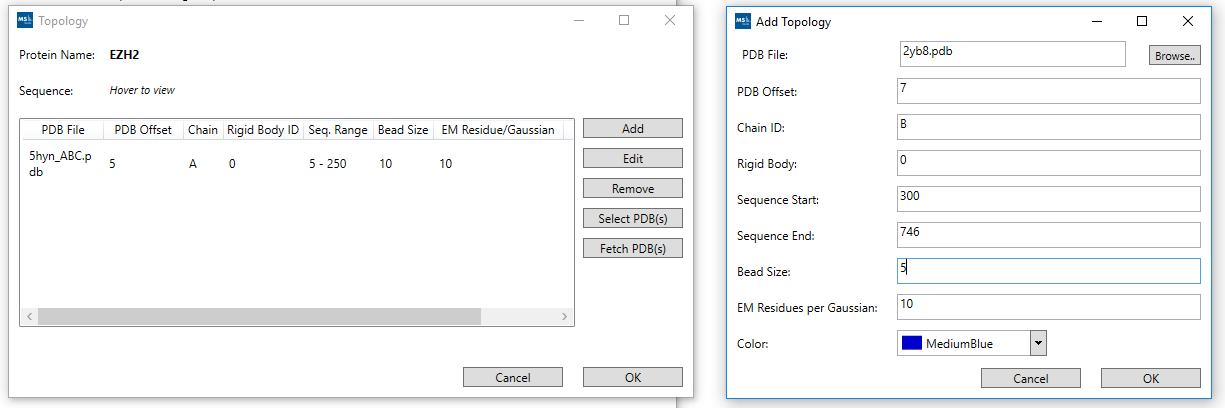
3. Add Proteins Topology:
The Topology record can be edited to set the start and end of the sequence together with the PDB Offset etc. Once you click Ok you will be returned to the Add Proteins wizard screen so that you can do the same for each of the Proteins involved. The representation can be adjusted e.g. two structures can be assigned to a single sequence and bead size can be adjusted. Once you have completed all the Proteins that you wish to amend. You can click the Next button (at the bottom right hand corner of the screen) which will take you to the Add Link Data wizard screen.
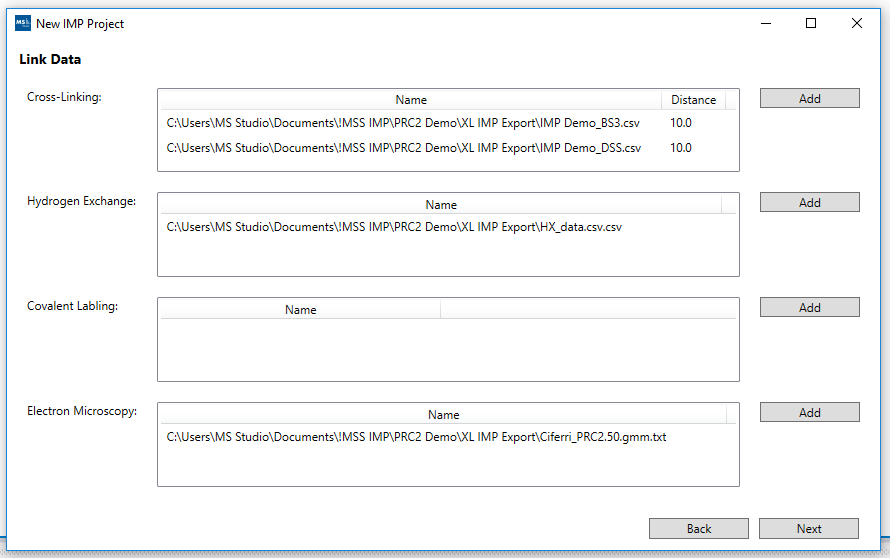
4. Add Link Data [XL, EM ...]:
Add Link Data wizard screen is where you can add additional data files including Cross-Linking, Hydrogen Exchange, Covalent Labeling and Electron Microscopy. These files will be included in their respective folders for the final output that is generated. Once you have completed your file selections you can click the Next button (at the bottom right hand corner of the screen). This will take you to the Configure IMP wizard screen.
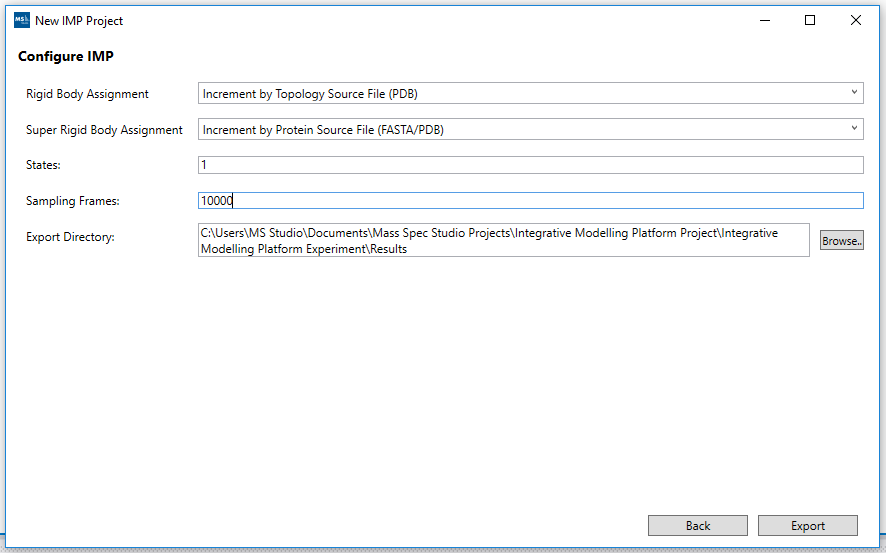
5. Configure IMP sampling frames:
The Configure IMP wizard screen is where we define the Directory path to export the data files and modeling scripts to. We also set the Sampling Frames and States here. The Ridgid Body and Super Ridgid Body assignments are available through the pick lists provided. The final step is to click the Export button (at the bottom right hand corner of the screen). This will produce the folder structure containing the Topology and YAML Config file together with the raw data files that you selected in the wizard steps ( data folder ). It also adds a folder with the modeling scripts needed (imp_model) to perform the job run using the python driver script provided.
6a. Exported directory - data files:
6b. Exported directory -python driver script:
Next: Deployment to HPC platform running IMP package :
We make use of a setup script from github gist in order to provide the commands needed to get the sample project from github. This brings with it the example files and scripts that we will be using to complete this demonstrating.
#### get the setup script from github gist and review before running:
curl -LOk https://gist.githubusercontent.com/pellst/4853822ea5ca74785af61d0ad39cf84d/raw/uoc_mss_prep_step1.sh
chmod 755 uoc_mss_prep_step1.sh
#### run the script uoc_mss_prep_step1.sh in order to get the sample folders and scripts setup ./uoc_mss_prep_step1.sh
#### in the folder /scratch/$USER/imp/imp_msstudio_init-master/mss_out/imp_model, the following shell scripts are now available
uoc_mss_prep_step1.sh
uoc_mss_prep_step2.sh
uoc_mss_prep_step3.sh