Crosslinking Mass Spectrometry (CRIMP)
The crosslinking package is a collection of tools built on top of the Mass Spec Studio core framework for processing and visualizing crosslinking mass spectrometry data. Crosslinked peptides as well as other variations such as free peptides and dead ends are discovered using a probablistic data reduction search algorithm. The final search results can be inspected and refined manually in the validation viewer.
Identification Algorithm
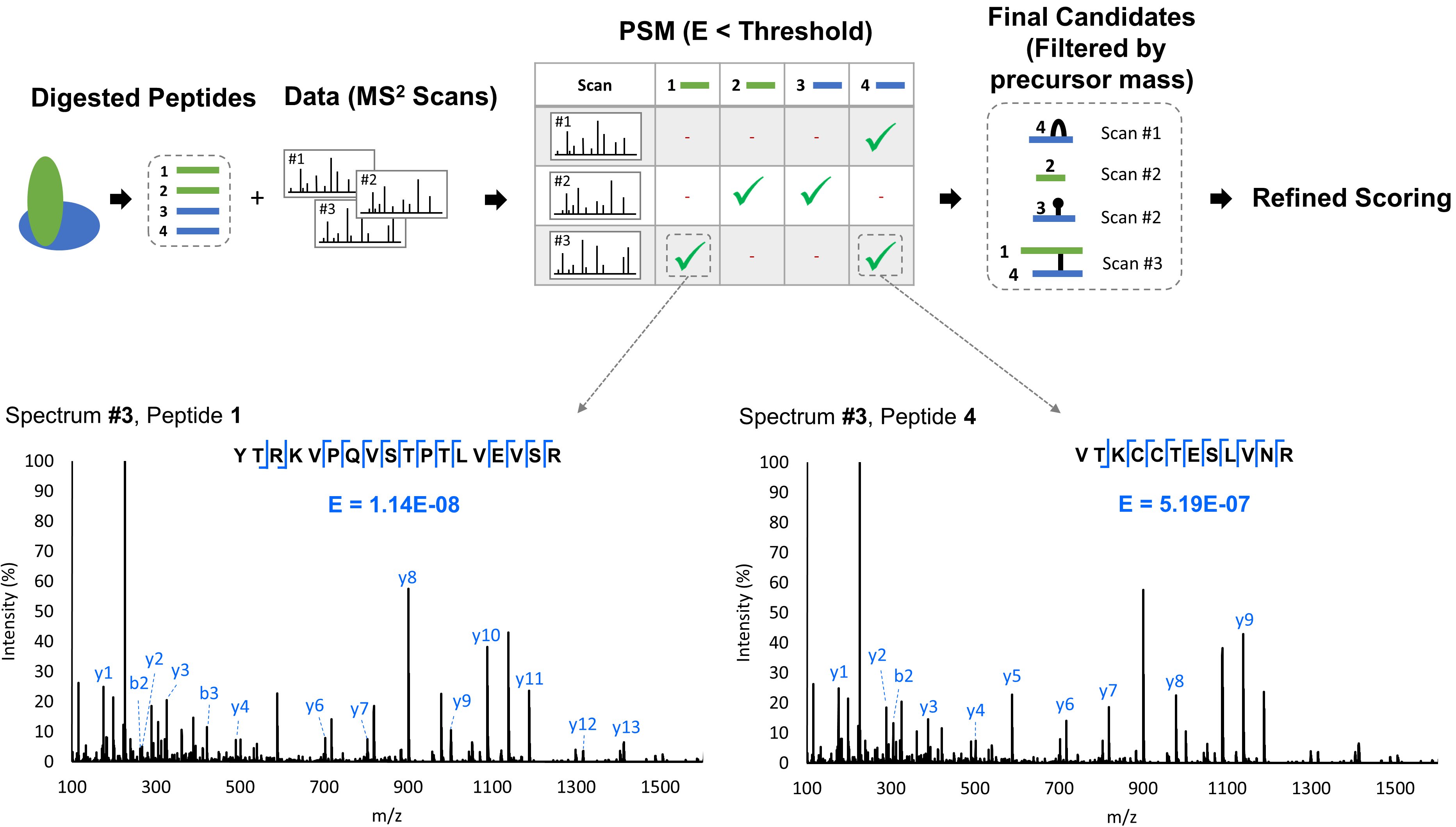
The probablistic data reduction strategy is designed to address the O(n2) time complexity problem associated with investigating all possible pairs of peptides involved in crosslinking. Crosslinked peptide pairs are generated only for peptides which have evidence of fragment masses in the same MS2 spectrum. To achieve this, expectation values (E values) are calculated for each peptide-spectrum match (PSM) using a variation of the Open Mass Spectrometry Search Algorithm (OMSSA). Only peptides with acceptable E scores in the same MS2 spectrum are considered for the refined crosslink analysis:
In the refinement phase, all peptide types (crosslinked, self-linked, dead-end) are re-scored using all of the available fragment data, such as crosslinked or modified fragments to generate the refined E values. The final scores -ln(E) are compared with the decoy database scores to calculate a cutoff score based on the False Discovery Rate (FDR).
Note: The open-mass search option improves crosslink detection sensitivity during the data reduction stage by considering open-mass modifications in addition to free fragments to calculate the PSM E score.
Validation
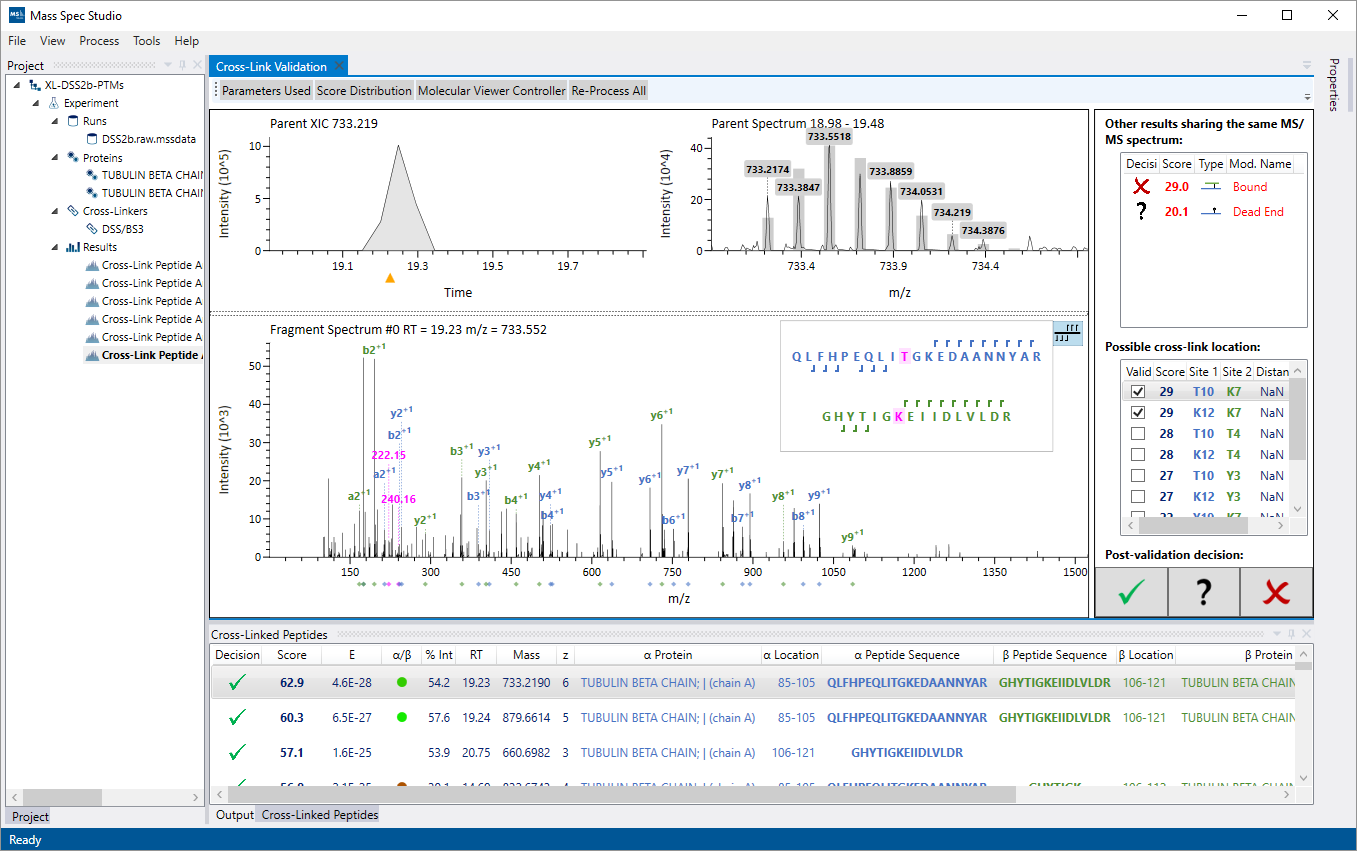
The final list of identified peptides can be inspected using the manual validation interface. A user can explore the raw MS and MS2 data, view individual fragment assignments, select crosslink sites, and accept or reject an identification result.
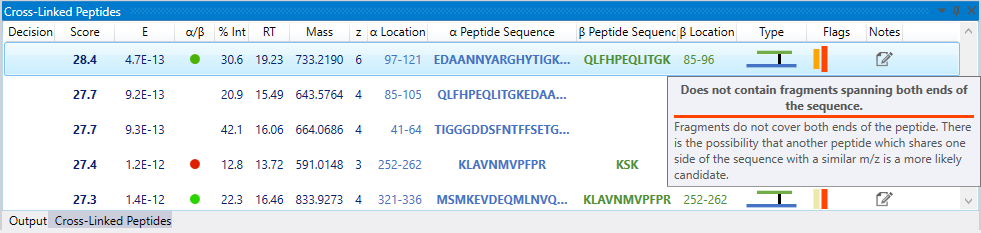
Warning flags inform a user of any potential problems with an identification. The flag color represents the severity of each problem, with more information available upon mouse hover-over.
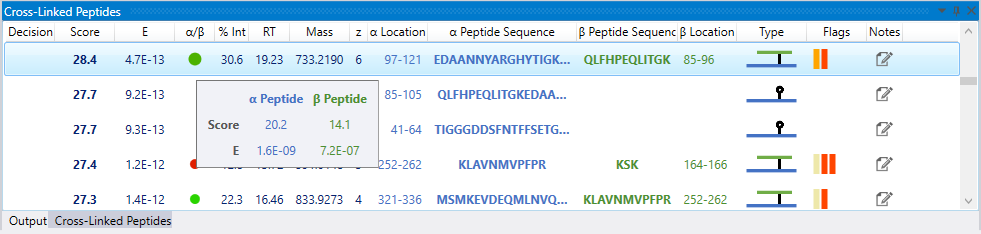
An additional flag which indicates the quality of a crosslink identification is the α/β flag. It represents the individual E score ratio between the peptides involved in a crosslink. Large discrepancies are shown in red.
Reference
[Link]: Sarpe V, Rafiei A, Hepburn M, Ostan N, Schryvers AB, Schriemer DC. 2016. High Sensitivity Crosslink Detection Couple With Integrative Structure Modeling in the Mass Spec Studio. Mol Cell Proteomics, 15(9):3071-80.