HX-PIPE Tutorial
In this tutorial, we will explain how to:
More information and sample data
[Link, Data] Raval S, Sarpe V, Hepburn M, Crowder DA, Zhang T, Viner R, Schriemer DC, Improving Spectral Validation Rates in Hydrogen-Deuterium Exchange Data Analysis. Anal Chem, 93(9):4246-4254(2021)
Create Project
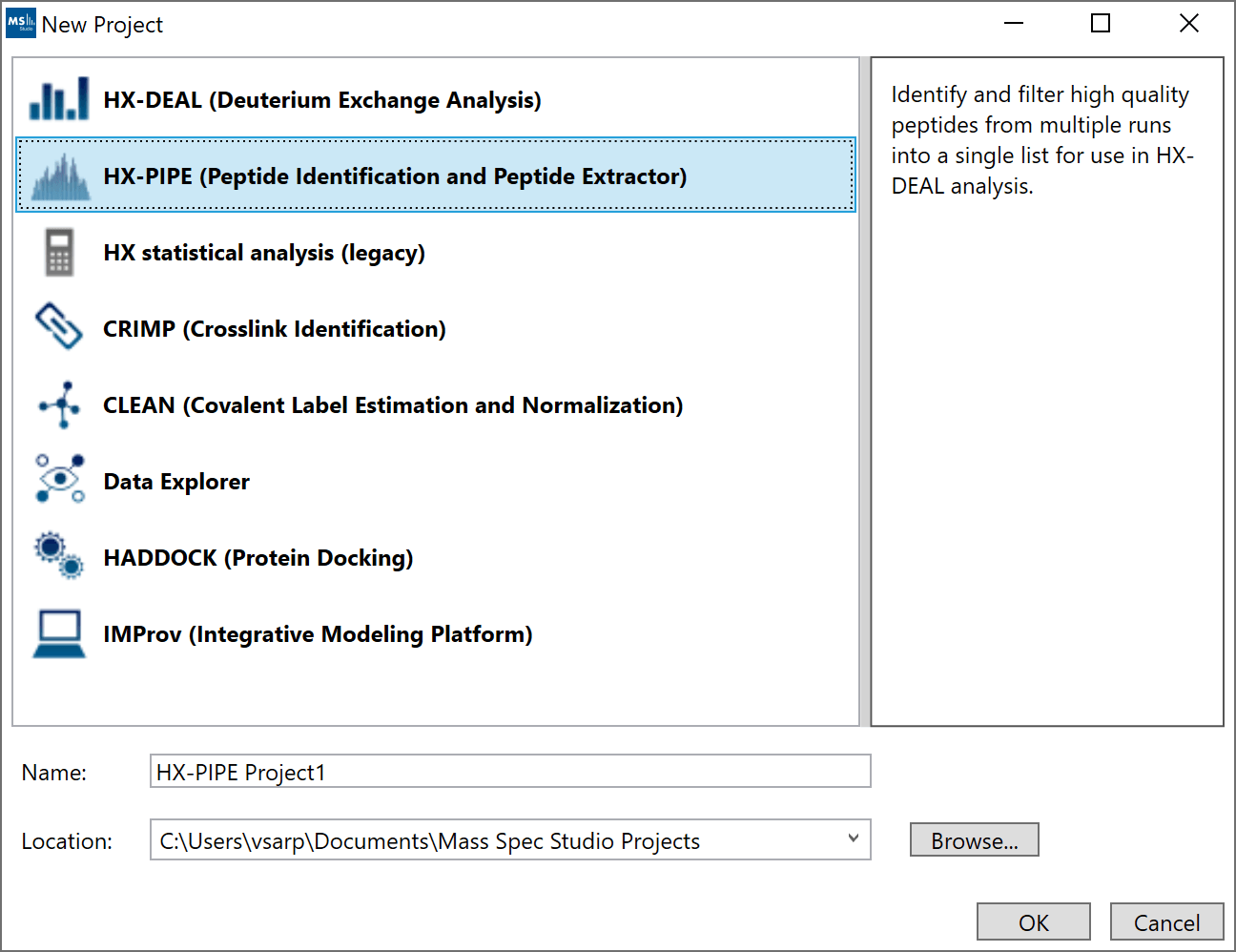
1. “New Project”: Create a new HX-PIPE project by selecting “New Project” and selecting “HX-PIPE. The default location for new projects will be inside your home directory. You can select a different location by clicking “Browse” and selecting a new directory.
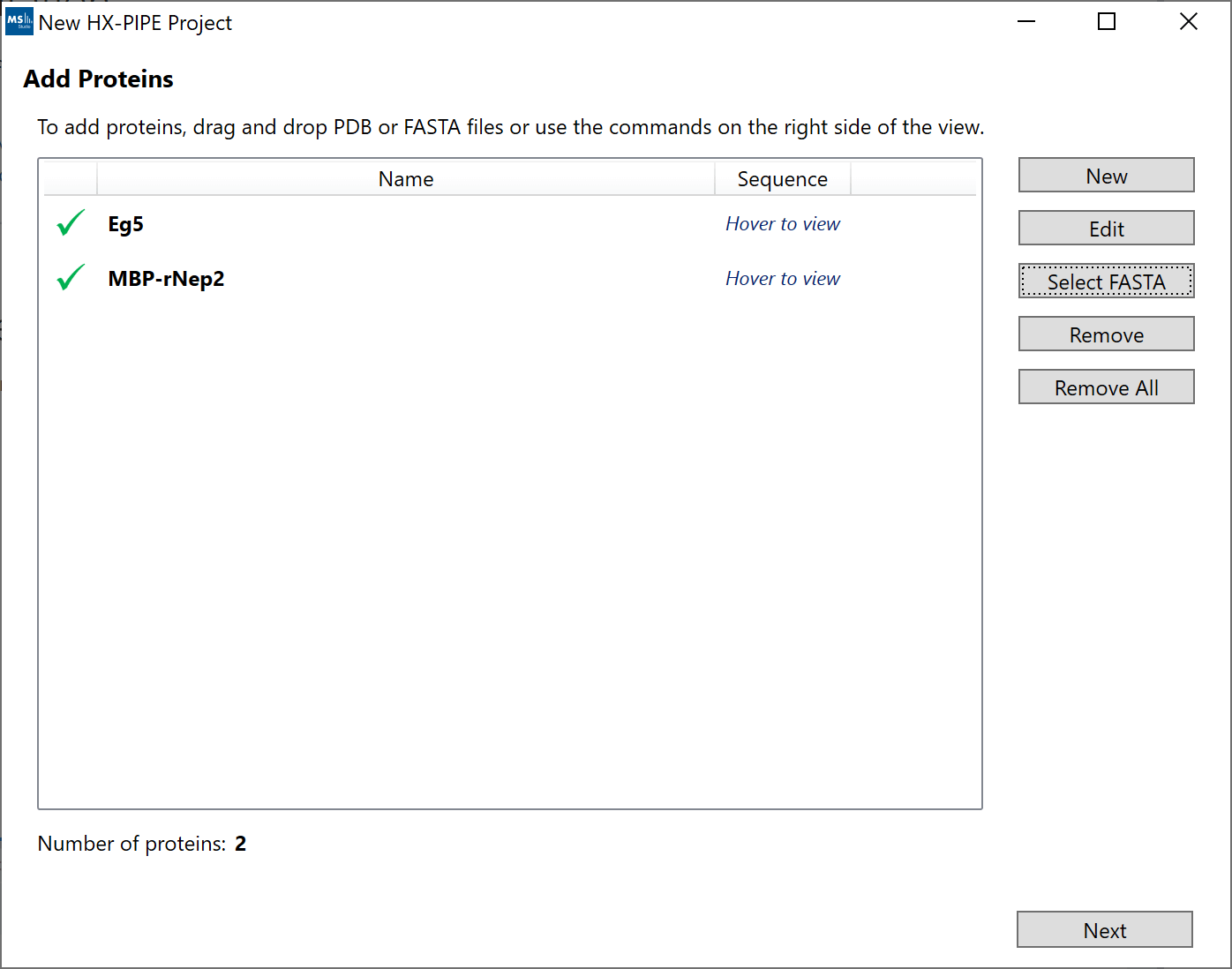
2. “Add Proteins”: If you have more than a single protein or wish to use sequence visualizations, you can add your protein sequences either manually (“New”), via a FASTA file (“Select FASTA”). Each protein will be listed as a separate row.
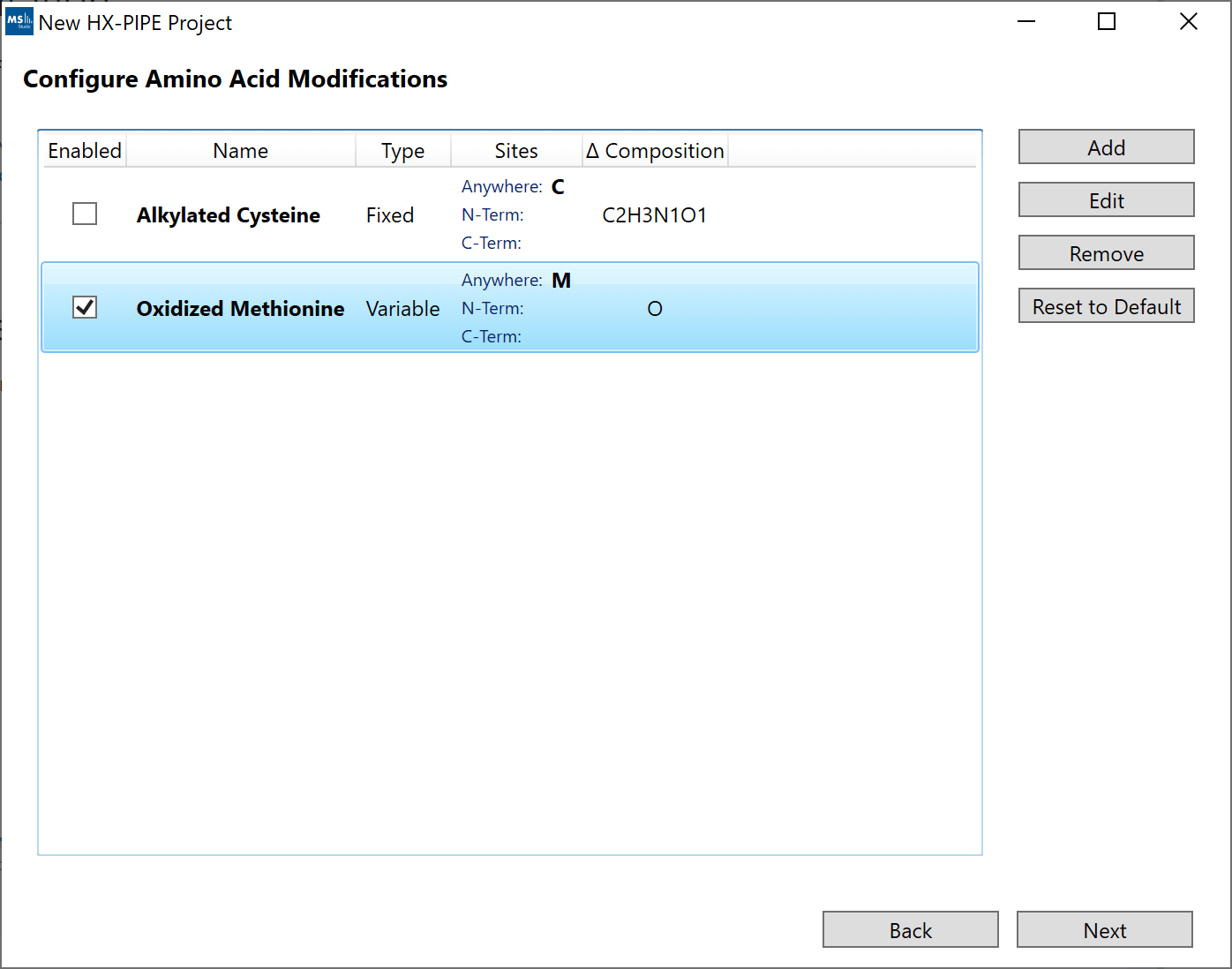
3. “Configure Amino Acid Modifications”: Post-translational modifications can be defined here. Click "Add" to define a new modification, these will be saved and available for future projects.
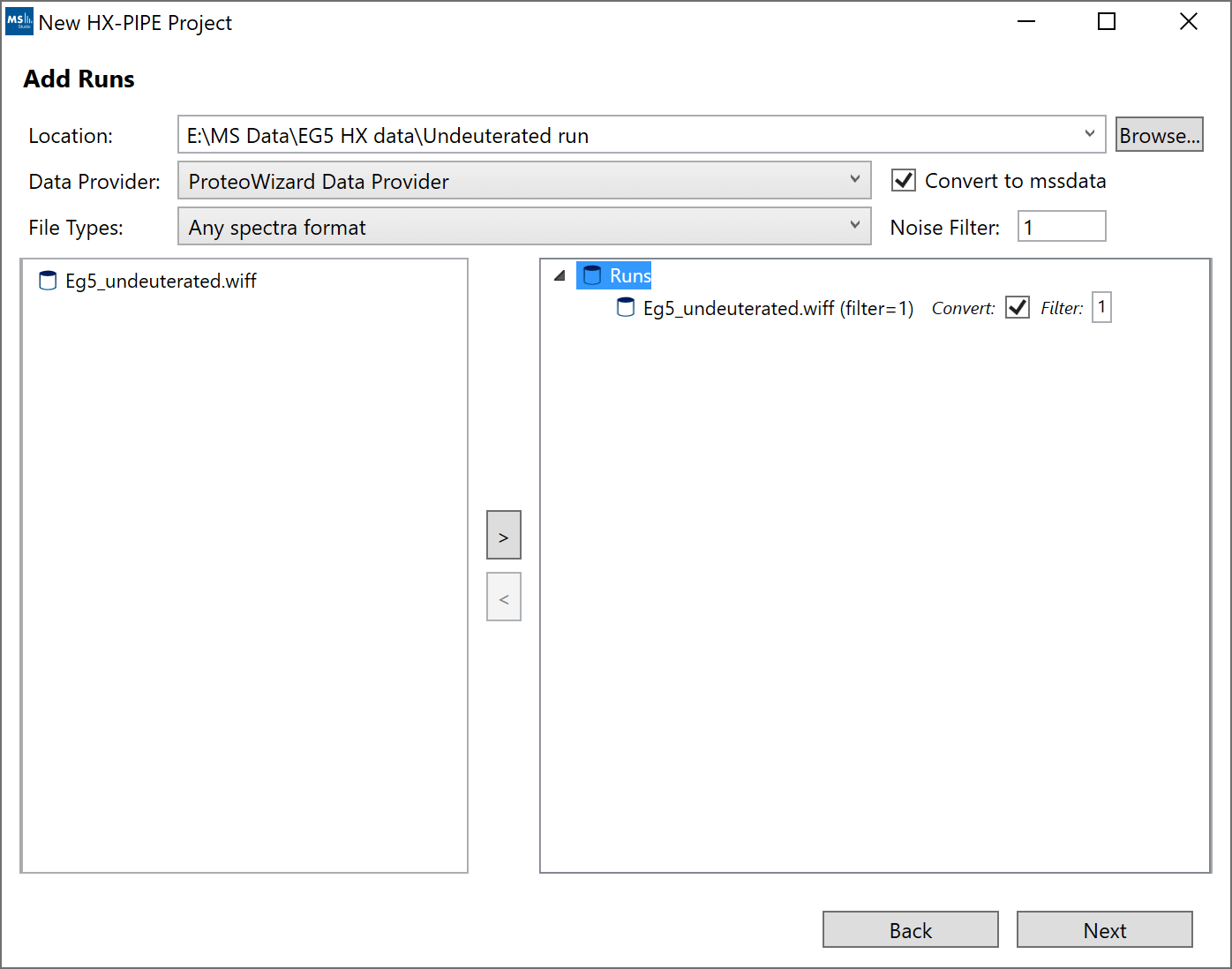
4. “Add Runs”: Follow these steps to add your data:
- a. Click “Browse” and select the root folder which contains your raw data files. In cases where the data files themselves are directories (example: Waters, Bruker), please make sure the containing folder is selected not the top-level .raw or .d directory itself. Note: Not all files have to be located under the same root folder from the start. After you add some files, you can still “Browse” to a different root directory and select additional files from the new location.
- b. For most raw vendor files, we recommend to use “ProteoWizard Data Provider”. If you have Mass Spec Studio converted files (.mssdata, .mssmeta), select “Mass Spec Studio Data Provider”.
- c. We recommend you enable the “Convert to mssdata” checkbox. When ON, raw vendor files will be converted to the .mssdata format which enables super-fast searching at the cost of some additional disk space for the .mssdata files. You can still proceed without converting the files, but the processing will be very slow. The “Noise filter” value is a multiplier of the minimum signal in each spectrum. A noise filter of “2” will remove any intensities smaller than 2 * mininum. A value of “0” will not remove any data.
- d. Select your replicates (shift+click or ctrl+click for multi-select), select the appropriate Protein State/Labeling node and click the “>” button. If you make mistakes, you can remove runs using the “<” button.
Note: The raw files will also be converted to mzXML during project creation. This is required for external search engines (MS-GF+). If you using Mass Spec Studio Data Provider, please ensure that either: (1) mzXML file already exist in the original repository for each run or (2) pre-convert runs to mzXML (MS2 scans are required).
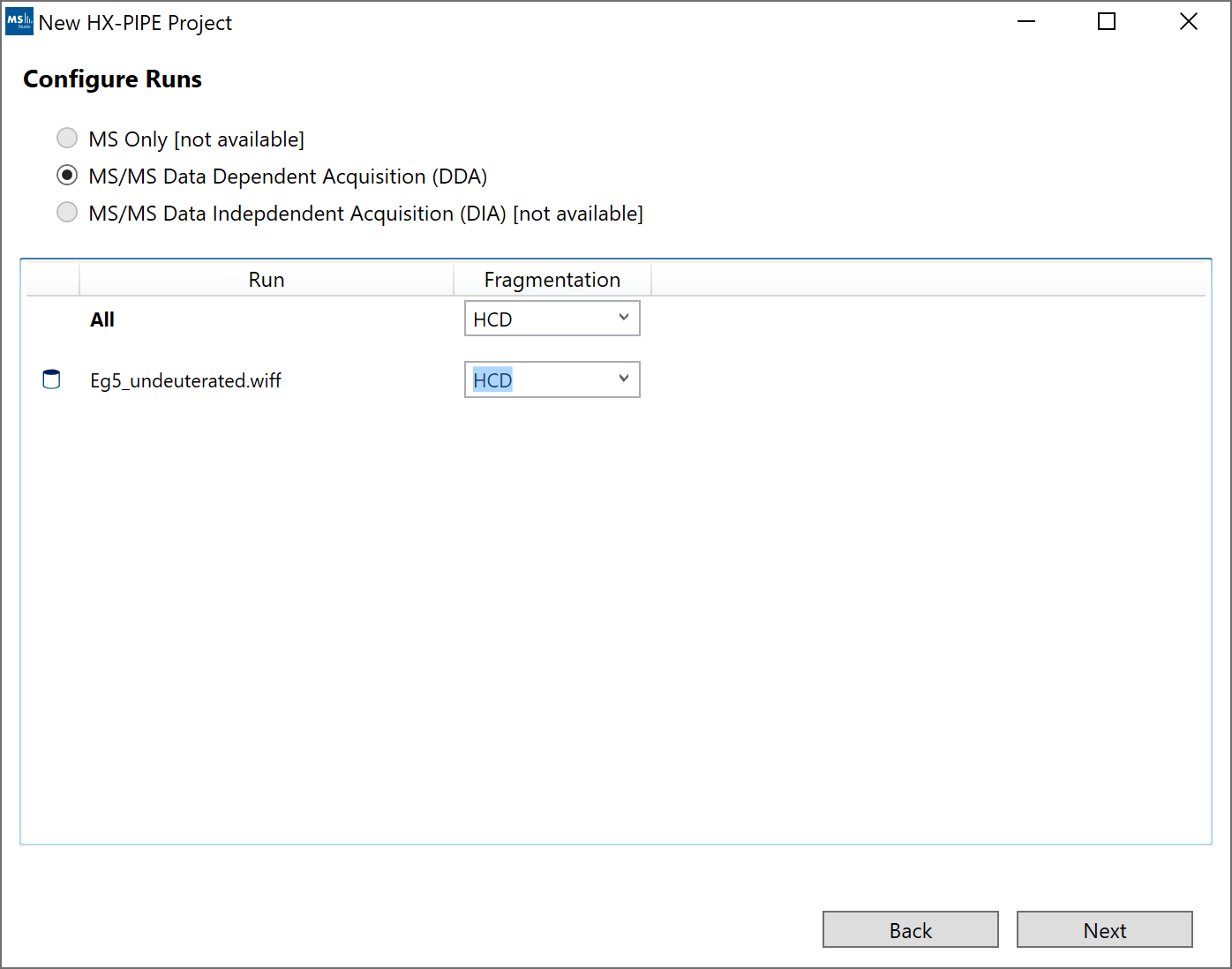
5. “Configure Runs”: For most files, select the fragmentation method of the MS2 spectra. For Waters files, you will have the option to change the default lockmass settings. These settings will only be applied if the data is not already calibrated from ProteoWizard. The “Filter” settings will default to the last function in each data file. If your reference function is not the last function, you must provide the function number manually.
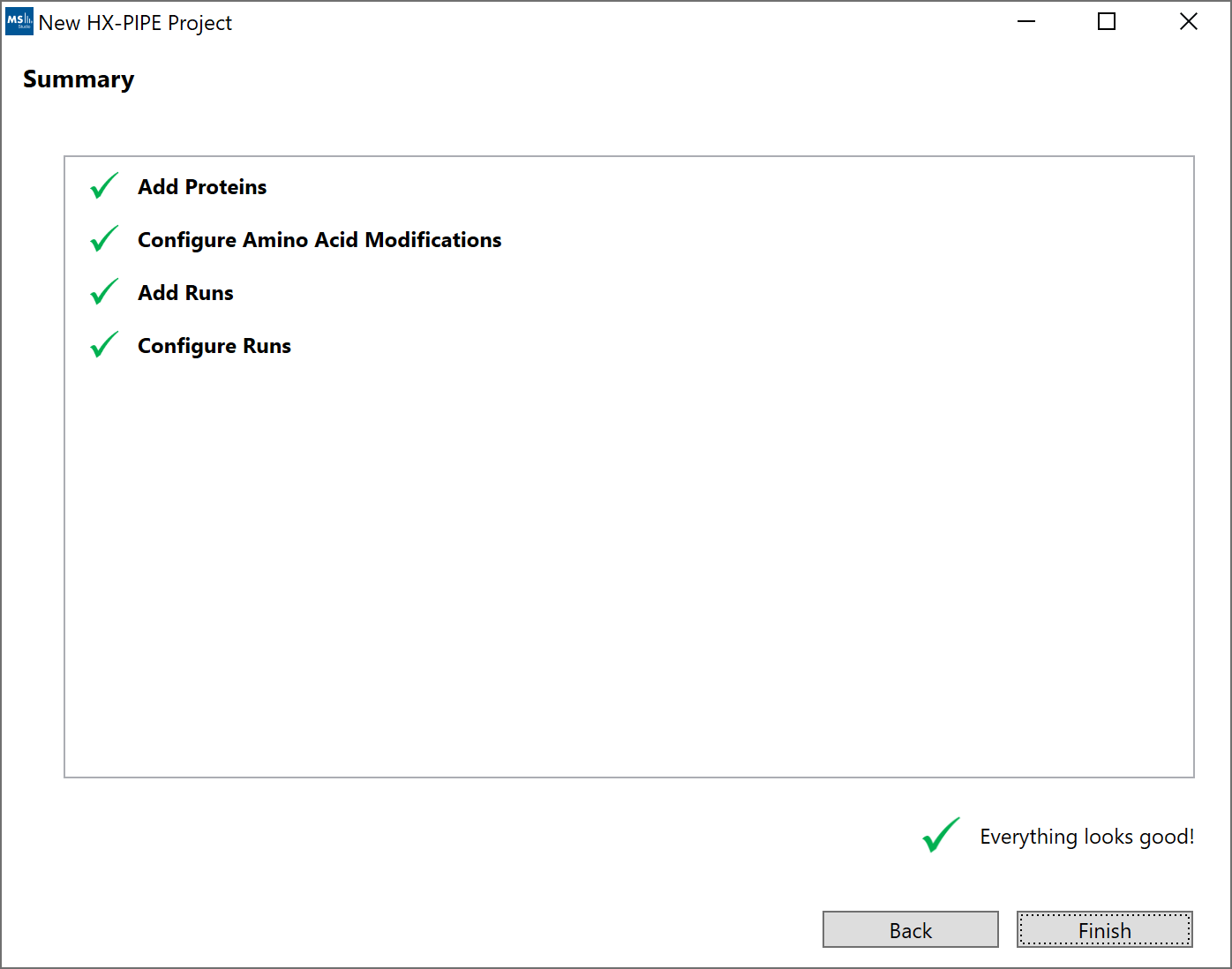
6. Summary”: The summary screen will show a top-down view of the project setup. Any missed errors or hints will appear under each step. Clicking on a row in the summary table will navigate back to that section of the wizard.
9. Complete the setup wizard by clicking “Finish”. A progress bar will appear to track the creation of the project:
Project Structure
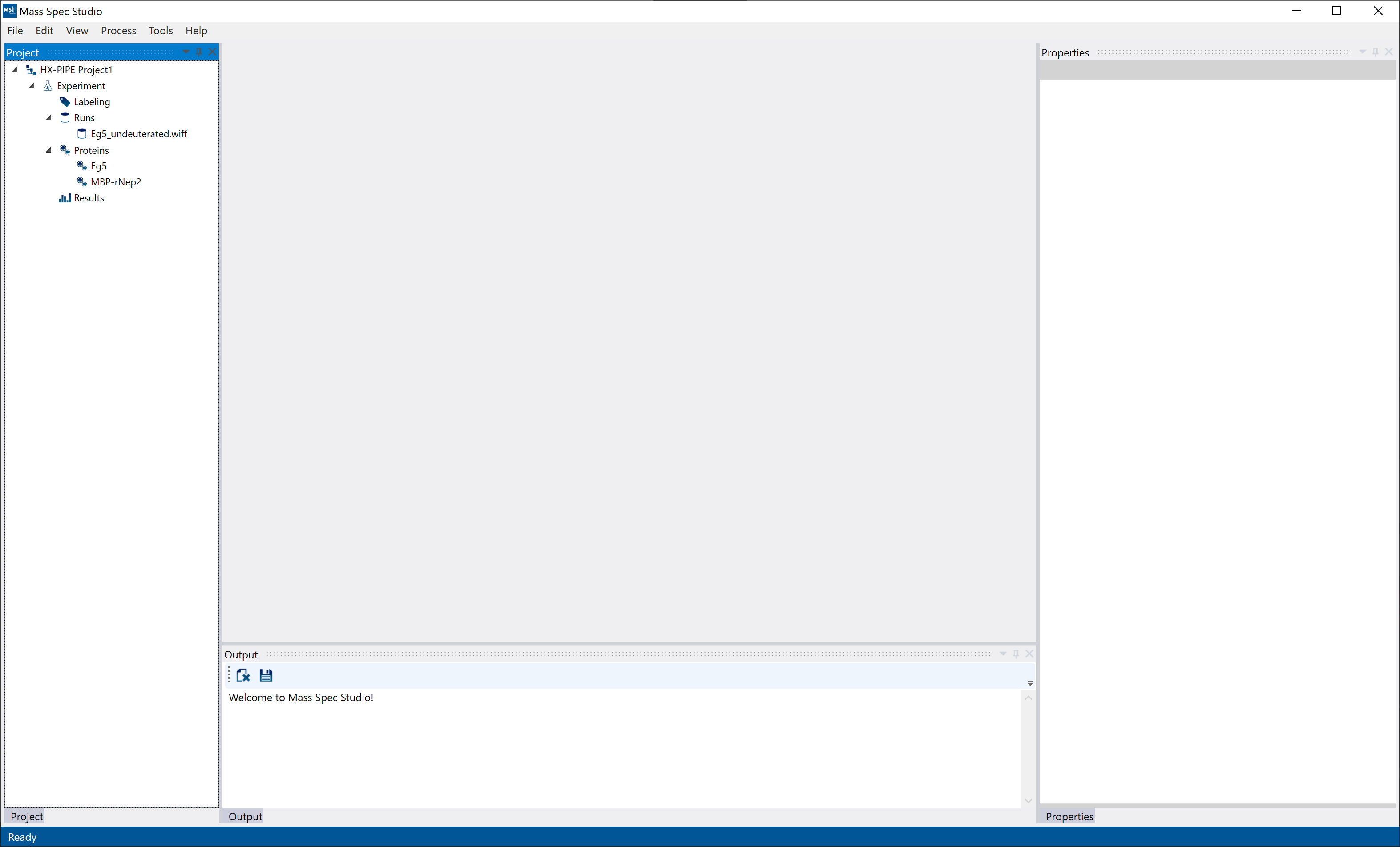
Once the HX-PIPE project is created, you will see an empty project view with an empty document section (middle) and left, bottom and right-side panels. The project tree structure will appear on the left panel with the runs, proteins, and results. The right panel will be a general “Properties” view where information from selected elements will appear. The bottom panel will initially contain the “Output” for an empty project. Once processed, the peptide list and feature list will also appear on the bottom panel.
Hint: Panels and documents can be un-docked and moved to a different region or detached from the main window as a standalone window by clicking and dragging on the blue toolbar for panels and the title tab for documents. Clicking the pin icon will minimize a panel for a more compact view.
Hint: If any panels (left, right, bottom) are closed, they can be brought back from the “View” toolbar. To bring back the result views (middle - documents), you can double click on a result.
Processing
Once a project is created, you can open the processing window from the “Process” option on the main menu. The processing window has 3 areas:
- 1. The “Processing Routines” area will list the available routines that can be applied to your project. HX-PIPE has a single default “HX Peptide-Feature Identification” routine. Incompatible routines (for other project types) may appear in grey and can be hidden if the “Show Compatible Routines Only” checkbox is enabled.
- 2. The “Recent Parameters” area will save your most recent (last 5) parameter settings from previous successful runs.
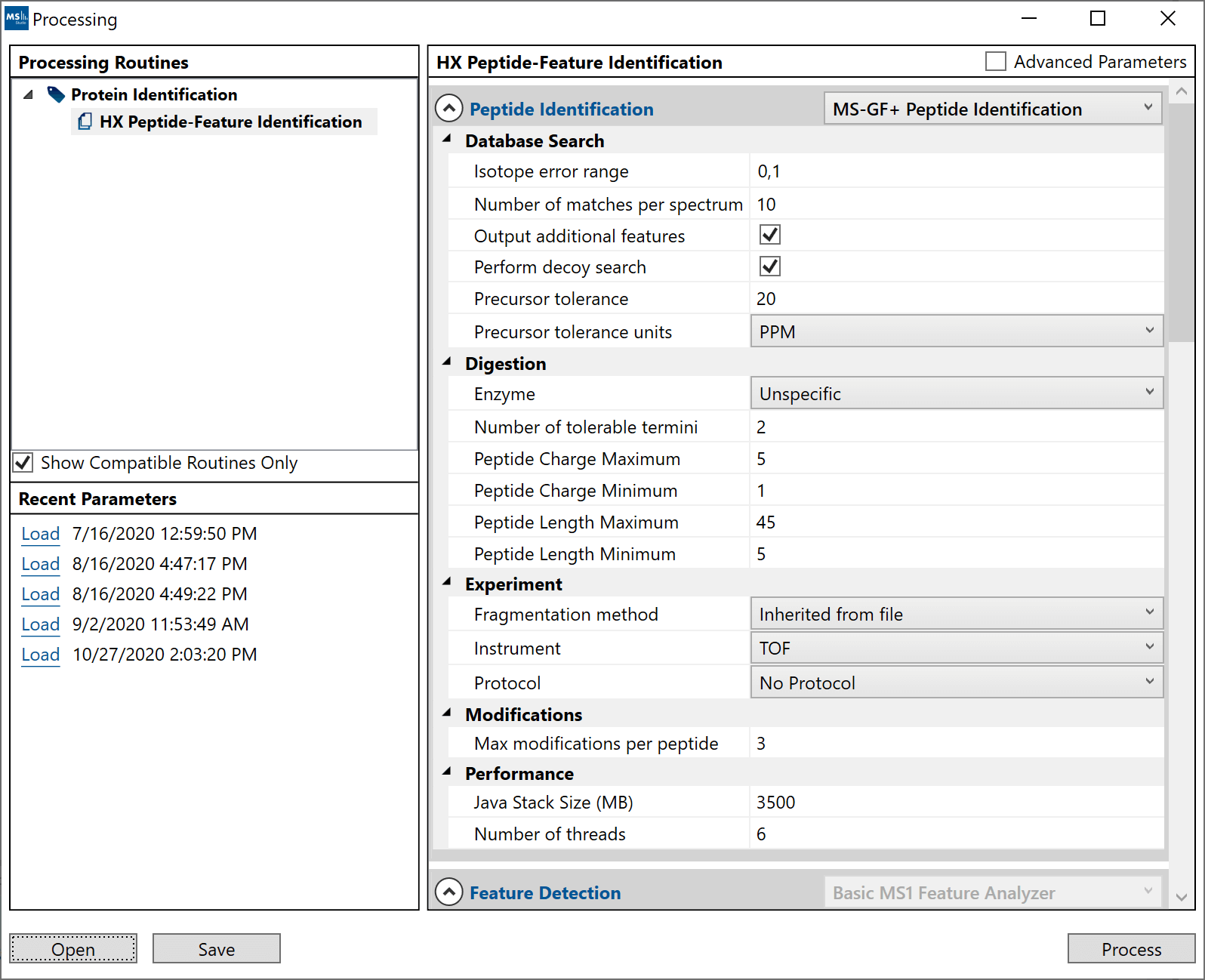
- 3. The main right-side area will contain the list of adjustable parameters for the selected processing routine. Advanced users can choose to display additional “Advanced Parameters” from the checkbox on the top-right, which enable more control over the analysis. Parameters can be saved/loaded by interacting with the “Recent Parameters” list or to/from file (XML) via the “Open” and “Save” button on the bottom left./li>
The search parameters for HX Peptide-Feature Identification:
- “Peptide Identification”: The peptide identification step is a
proteomic-style MS2 search to generate peptide-spectrum matches. This step has 2
options:
- “OMSSA+”: Uses a modified OMSSA algorithm with a user-defined E value cutoff.
- “MS-GF+: Uses the MS-GF+ algorithm. This option will be faster for data with lots of post-transltional modifications. It also requires mzXML (at least MS2) converted runs to exist in the same folder.
- “Feature Detection”: Settings for identifying MS1 features.
- “Feature Intensity Threshold”: The absolute intensity cutoff across all MS1 scans. Peaks lower than the defined intensity are not considered for feature identification.
- “p-Value Threshold: The goodness-of-fit to a model-based isotopic distribution of the same mass. The model used is a Poisson model to generate the peak intensities and can be changed to an Averagine model if "Advanced Parameters" are enabled under the "Isotopic Profile Model" parameter.
- “Peptide Elution Time”: Settings for dealing with XIC data. The "Width" is used for aggregating MS1 data for peptide and features. The "Tolerance" is used for grouping similar features appearing at different RTs.
- “XIC Extraction”: The m/z window used in extract the XIC for a given peptide. The mass range will be applied around the best peak or the monoisotopic peak for XIC extraction.
- “XIC Smoothing”: The type of smoothing for the XIC.
- “XIC Peak Picking”: The settings for finding peaks in the XIC data.
- “MS Mass Accuracy: The m/z tolerance for selecting MS1 peaks and extracing chromatograms.
- “MS Extraction”: The ranges for extracting MS data once the best RT range for the peptide is found. By default, it will extract a “Summed” spectrum using the defined RT range. The “m/z Padding” is the m/z range relative to the monoisotopic.
- “MS Smoothing”: The type of smoothing for the aggregated MS data.
- “MS Peak Picking”: The settings for finding peaks in the aggregated MS data.
- “MS/MS Mass Accuracy: The m/z tolerance for selecting MS2 peaks and extracing chromatograms.
- “MS/MS Smoothing”: The type of smoothing for the aggregated MS2 data.
- “MS/MS Peak Picking”: The settings for finding peaks in the aggregated MS2 data.
The default parameters should work for most runs. If you are seeing instances where good peaks are not selected by default, try widening the “MS Mass Accuracy” and “XIC Extraction” ranges.
The processing routine will calculate deuteration for all your peptides across all your data file. When finished, a new result will appear in the “Results” section of your project tree. The processing routine can be re-run as many times as needed with different parameters combinations, with each result saved separately. Double-click on a result to open up the manual validation view as well as the rest of the visualizations in the middle document region.
Visualizations
There are several visuals for quick exploration of the protein/peptide/feature identifications once a result is generated in the previous step.
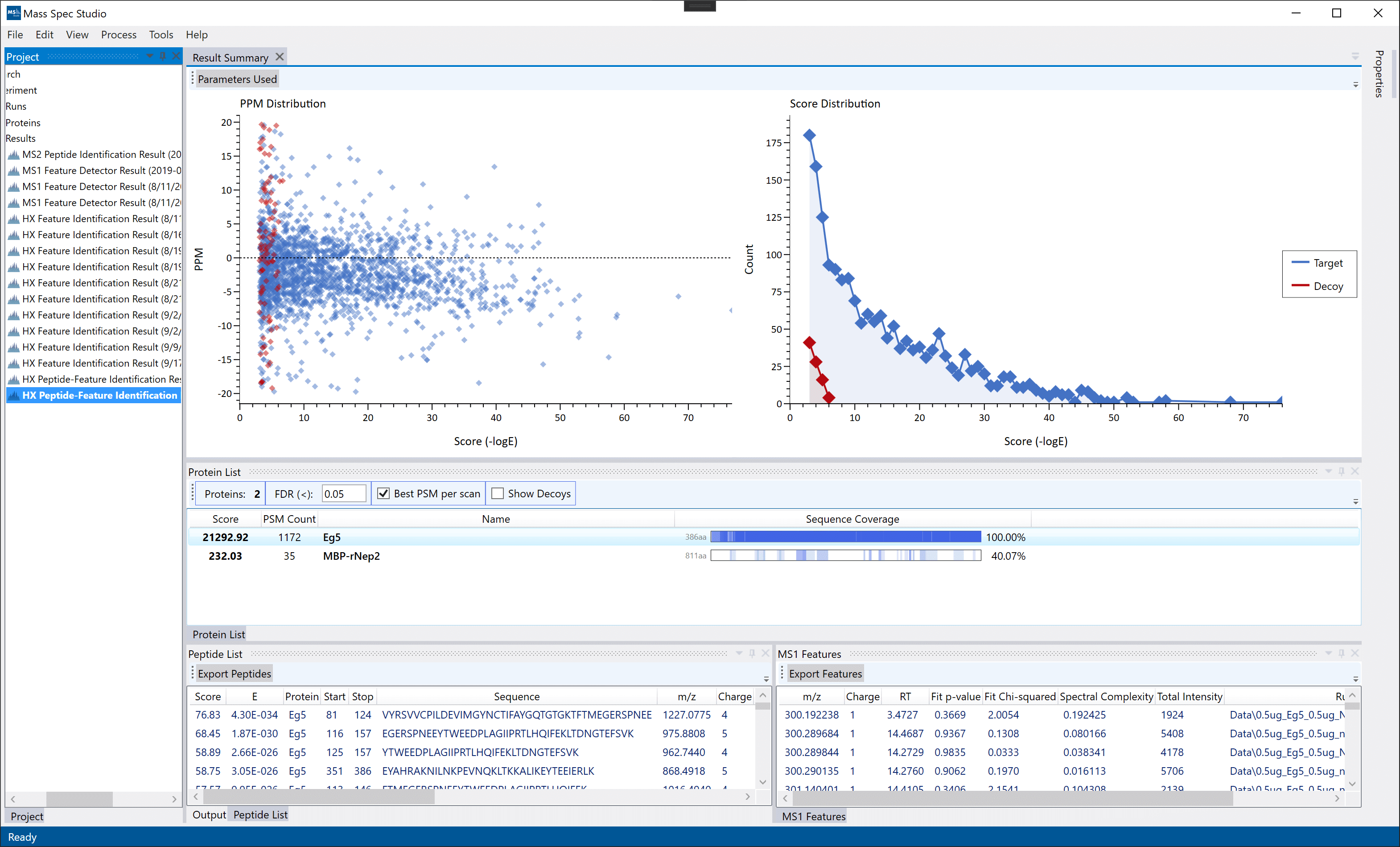
Result Summary: A top-down view of the overall search result. The two graphs display the results of the target-decoy result set in the MS2 search (decoy searches are less informative for really small datasets). The middle Proteins List table describes the protein identifications along with their underlying peptide-spectrum-match counts and scores. The bottom-left panel is a list of peptides from the MS2 identification step. The bottom-right panel is a list of features discovered by the MS1 feature finder.
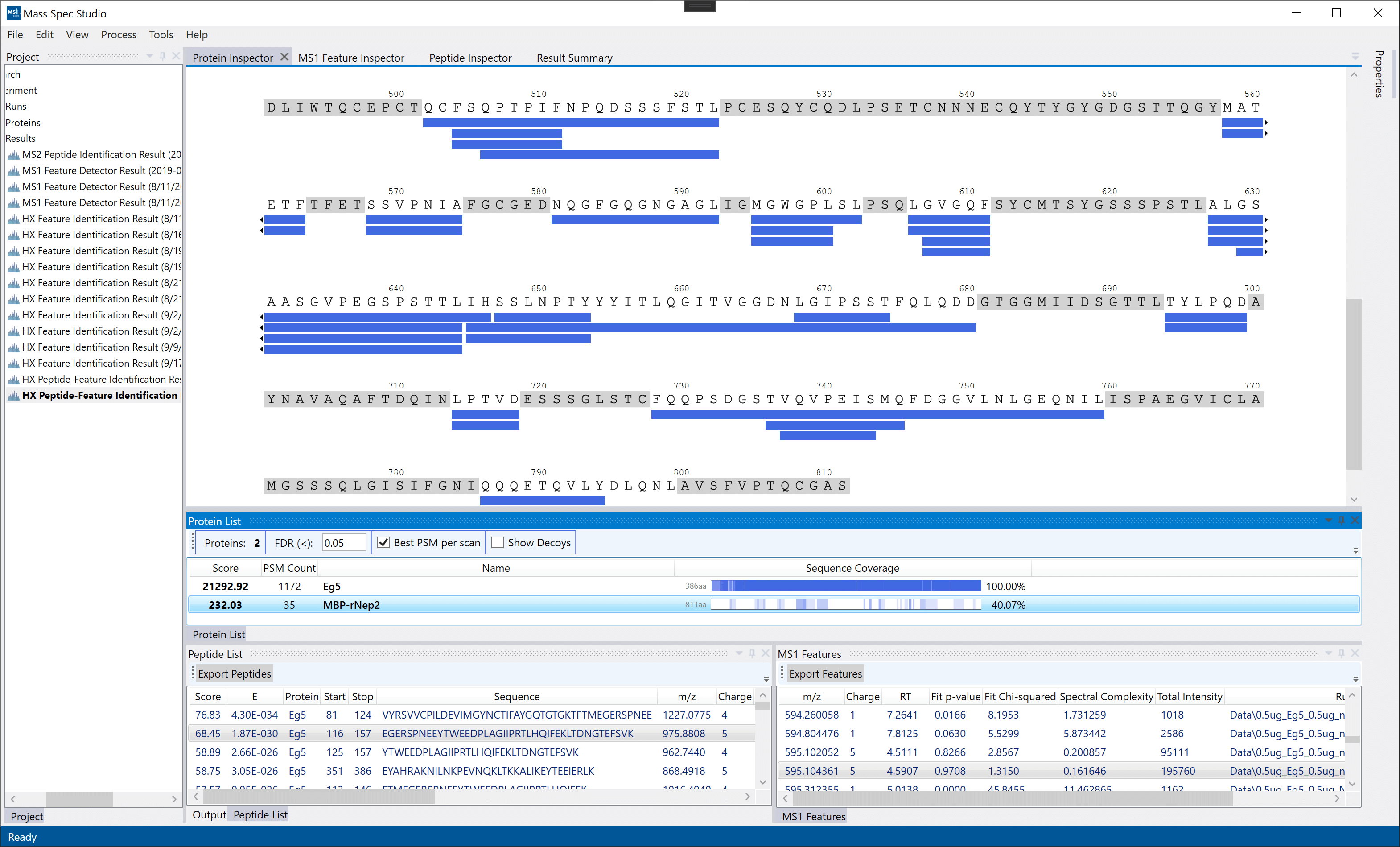
Protein Inspector: Displayed by double-clicking a protein result in the Protein List. This view maps all peptide identification from the MS2 search onto the protein sequence. The peptides are not yet grouped across runs (done during export), so repetitive sequences will appear multiple times. The "FDR", "Best PSM per scan" and "Show Decoys" options are meant to quickly inspect protein scores and will not affect the final peptide/feature results.
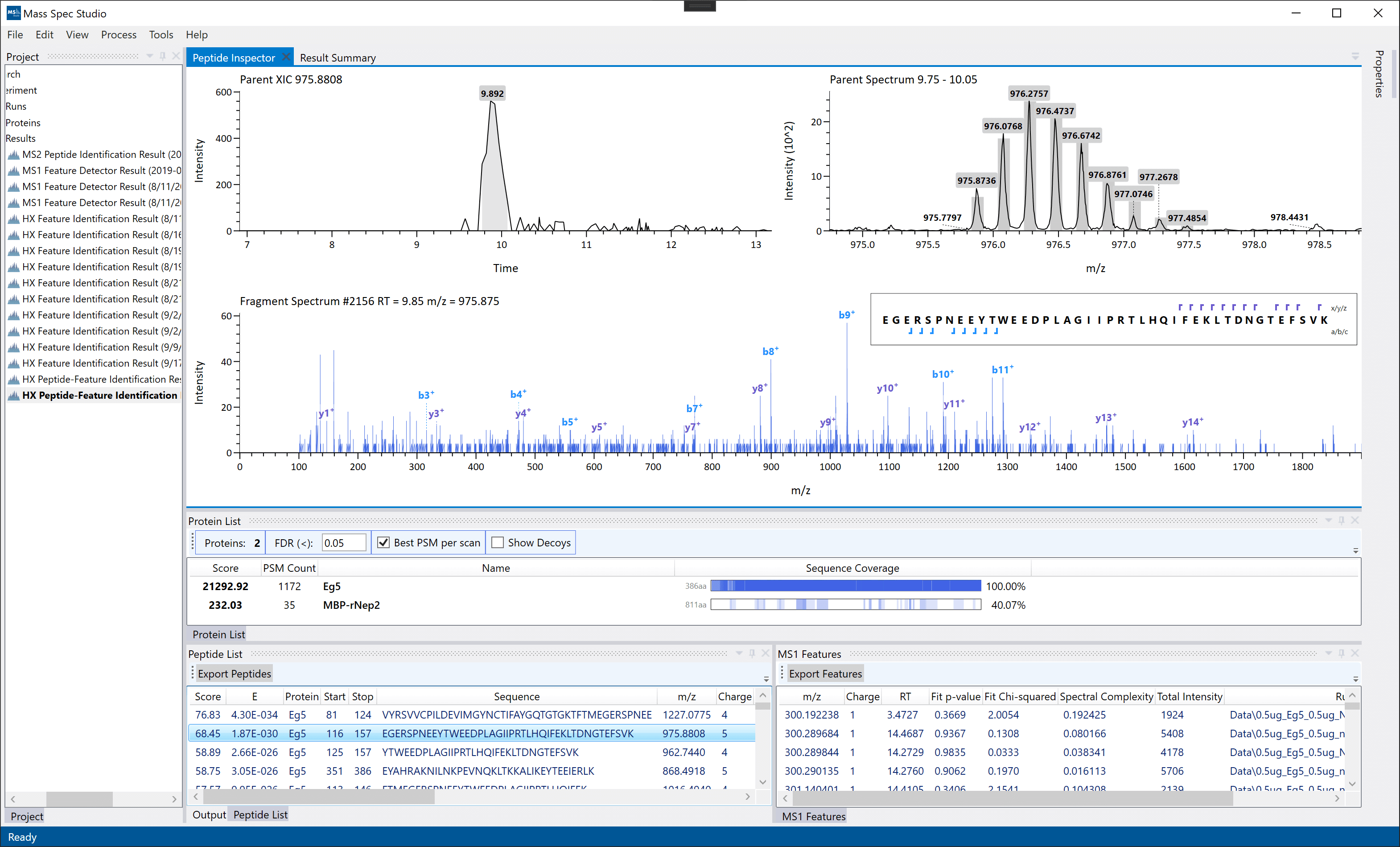
Peptide Inspector: Displayed by double-clicking a peptide identification in the Peptides List. This view displays the best MS2 spectrum (best PSM) of the peptide identification. Note: The fragment list may not match 100% of the fragments used during scoring, depending whether we have access to the internal scoring methods of the identification algorithm chosen (OMSSA+ or MS-GF+). These un-grouped peptides are exportable. Note: final grouping across between MS2 peptide identifications and MS1 features is done during export (File -> Export).
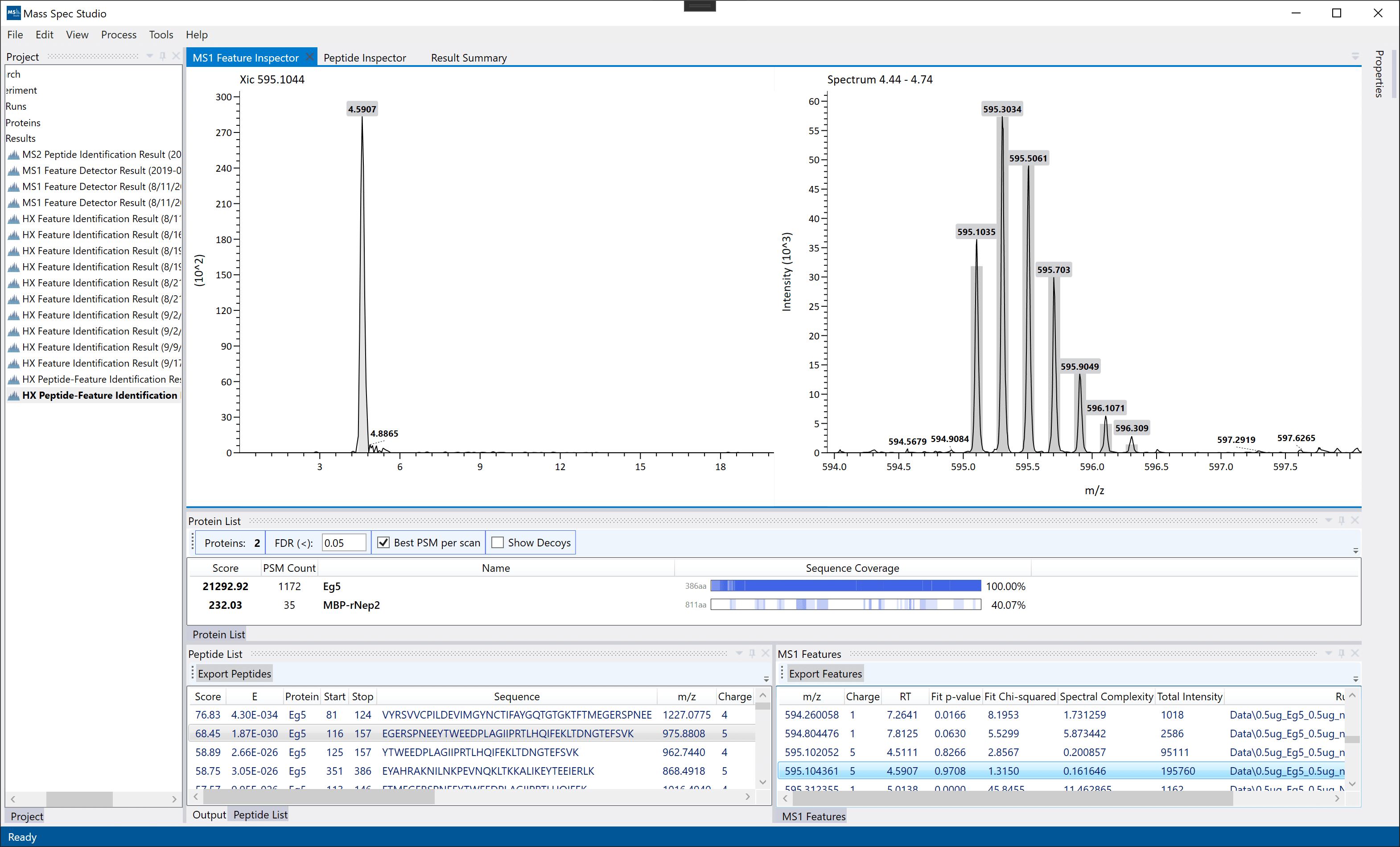
MS1 Feature Inspector: Displayed by double-clicking a peptide identification in the MS1 Features list. This view displays a selected MS1 feature. Similar to the Peptide Inpector, the results here are not yet grouped across all files. These un-grouped features are exportable. Note: final grouping across between MS2 peptide identifications and MS1 features is done during export (File -> Export).
Hint: Once all panels are activated, the user will have to manually navigate between them (not automatic). A useful method to be able to display everything at once is by using the dock/undock features (clicking/dragging the panel titles) and either floating them or docking them in a different configuration.
Export
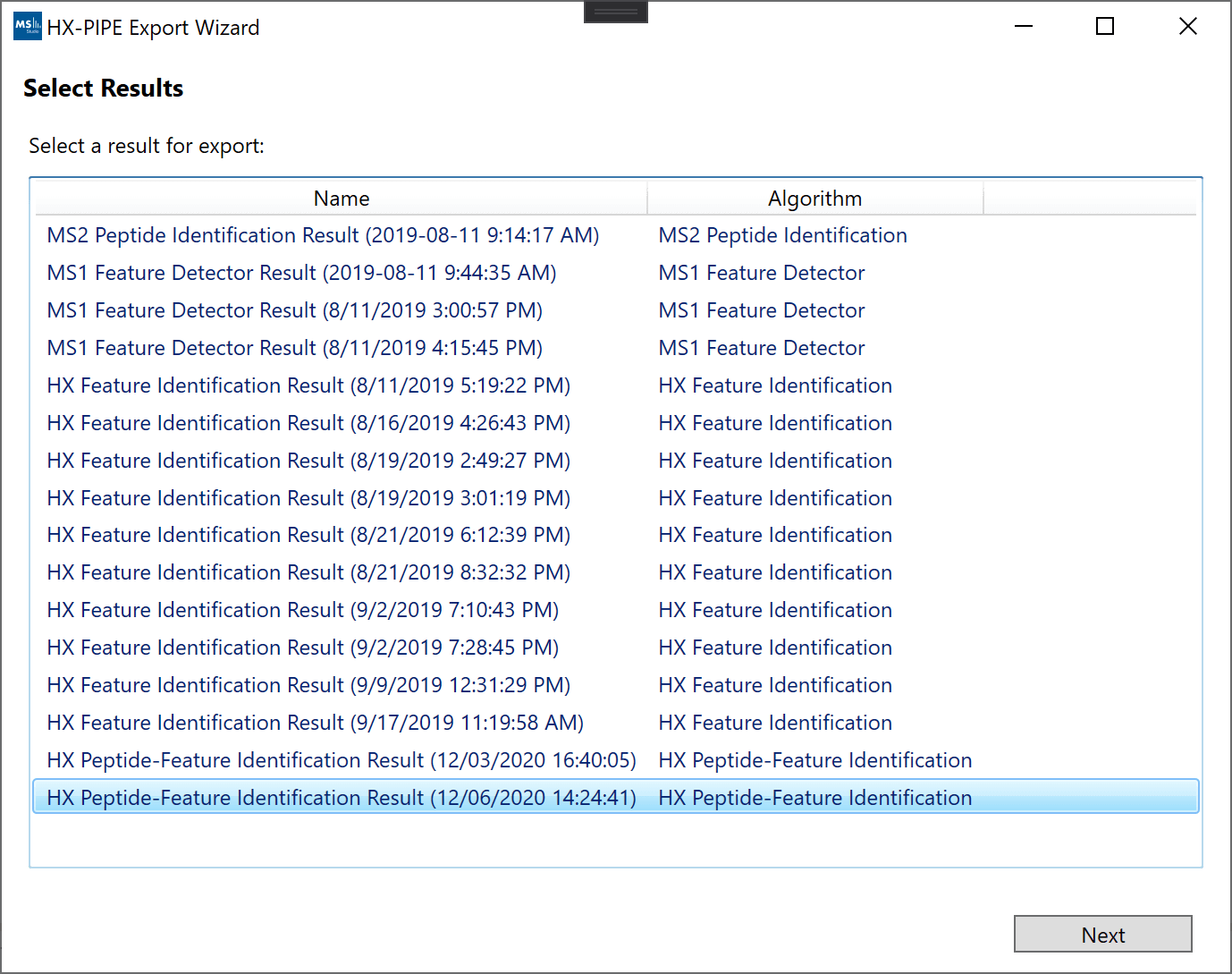
To export a result, you can click on “File -> Export” and use the Export wizard to select a result for final grouping and filtering.
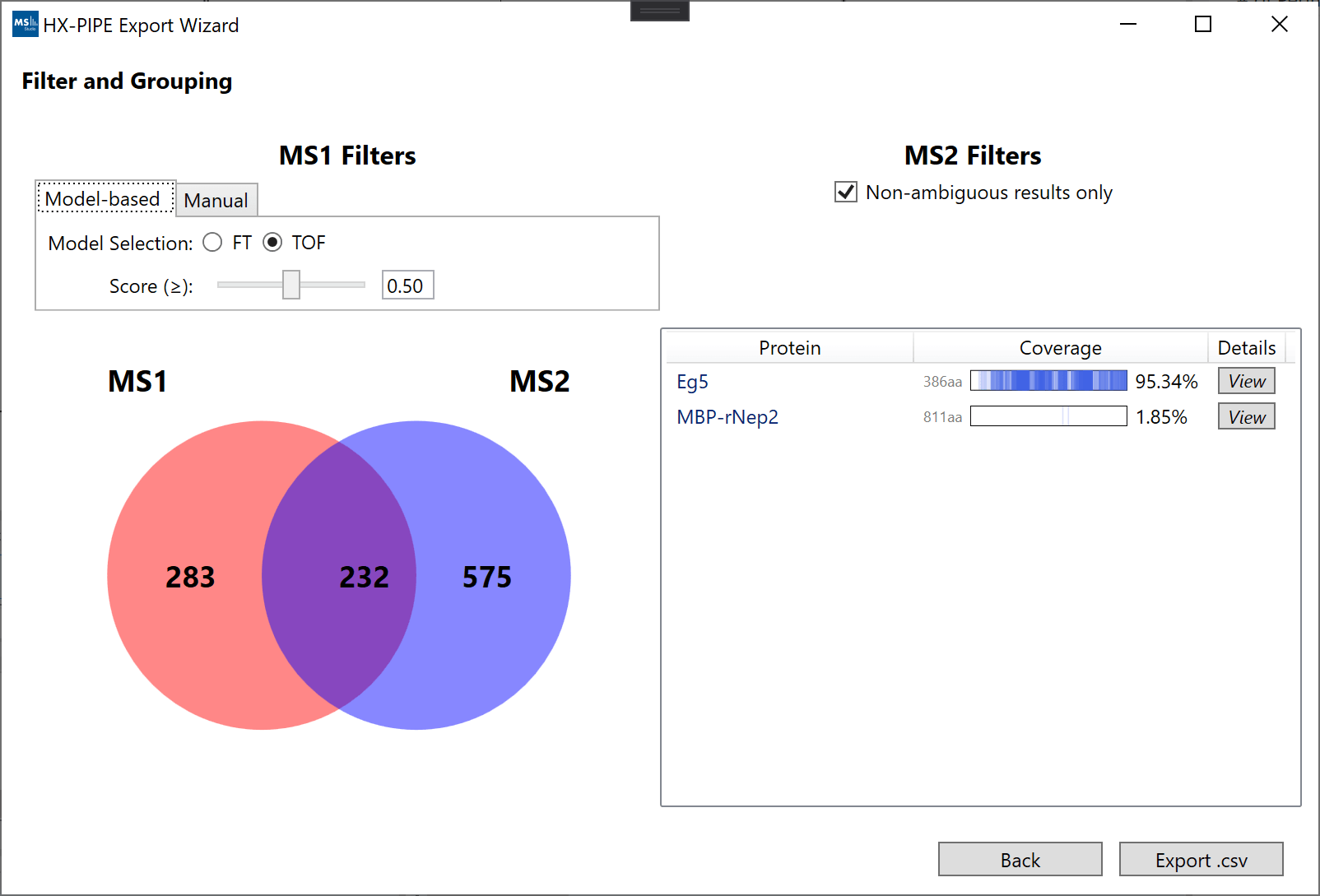
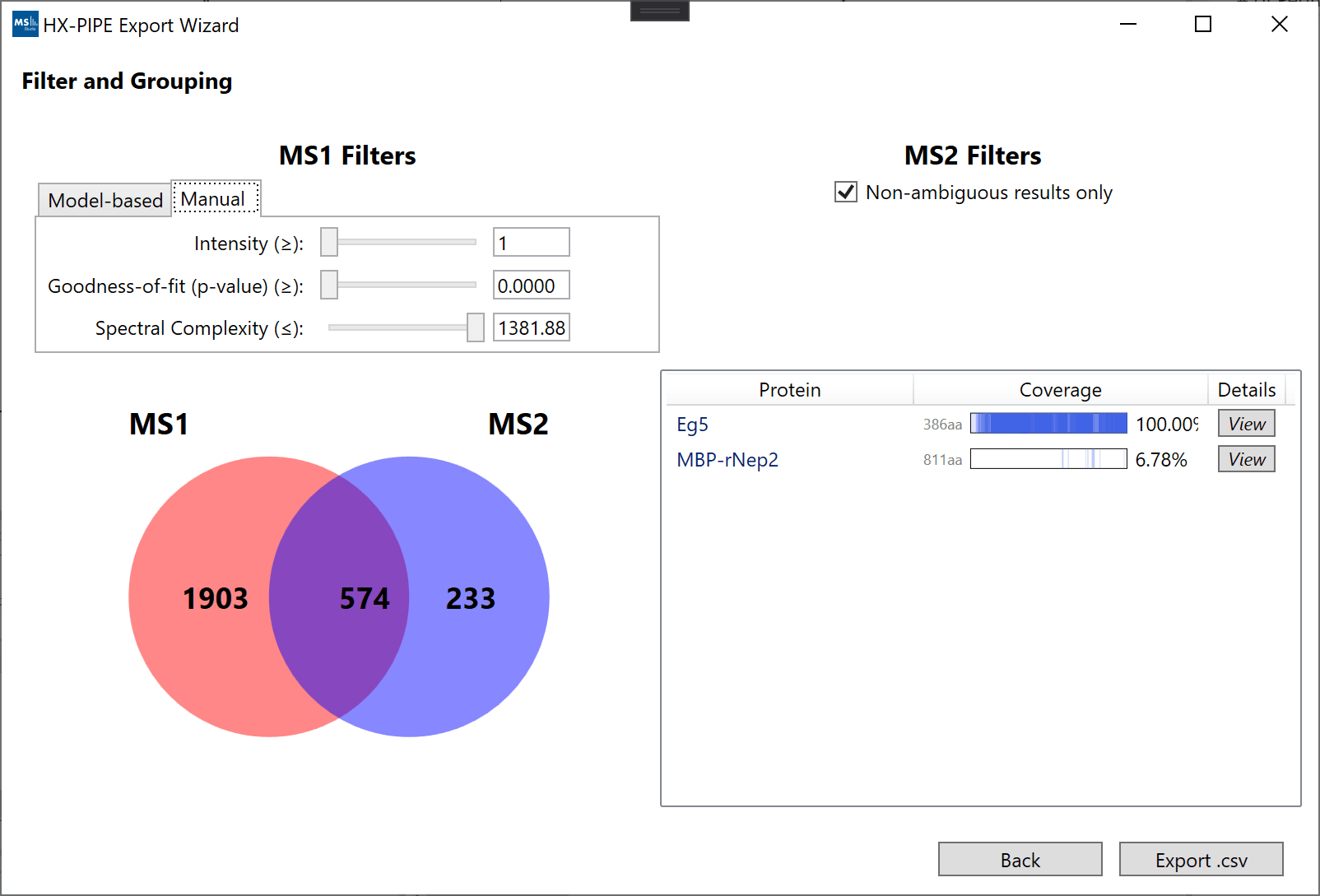
After selecting a result, you will be presented with the "Filter and Grouping" view. This view is split into 4 sections: (top left) the filters for MS1 features, (top right) the filters for MS2 identification, (bottom left) a Venn diagram of the final grouping/filtering results, (bottom right) a list of all proteins and their % coverage as a result of grouping/filtering.
The main interactive component is the MS1 filtering (top left) which reduces the number of MS1 features which can group with MS2 peptide identifications. The main idea is to achieve good protein coverage while only accepting a high-quality set of peptide identifications with high-quality MS1 data (overlapping region). We provide 2 methods to perform the filtering:
- “Model-based”: Uses a set of Logistic Regression models (one fit on Fourier-Transform data and one fit on Time-of-Flight) to provide a single simple cutoff based on 3 properties (goodness-of-fit, intensity, spectral complexity). Logistic regression models use 0.5 as the default cutoff, but the user can adjust this value. Higher cutoffs are more selective. The model type (FT or TOF) is usually automatically selected based on the original file extensions, but it can also be manually adjusted.
- “Manual”: Allows the user to set custom values for all 3 properties for filtering MS1 features (goodness-of-fit, intensity, spectral complexity).
A more detailed map of the coverage can be seen by hovering over the "View" button in the proteins table. Clicking the "View" button will open an interactive window with the detailed sequence map. Changing the filters will automatically update the coverage map.
After Export
The export results will be saved as 3 .csv files (common, ms1, ms2). The final high-quality peptide file to be used for downstream HX analysis (HX-DEAL) is "*.common.csv".
Note: For some large result sets, the export action may take anywhere between several seconds to a few minutes. During this time, the UI is disabled. Once the export is successful, the Export wizard will close automatically.